This is a story of how a small side project I started as an intrapreneur grew into a national information system. I am going to give the project’s full history, but if you wish, you can jump to the year 2009, when the system was born.
Diversity of Databases
Even though exhibitions are the most visible aspect of a natural history museum (think dinosaurs), the majority of work done in many such museums revolves around research. Museum scientists study biodiversity, relationships between organisms and changes in the environment. A considerable part of this research is based on scientific collections, which are larger than the public collections – by several orders of magnitude.
I have worked in different positions at the Finnish Museum of Natural History Luomus, the largest natural history museum in Finland, for over ten years. There are about 13 million specimens in Luomus’ collections, gathered from around the world over the last few centuries: insects, plants, parasites, lichens and so forth. The specimens also carry information about where they were collected and when. Without this information, the specimens would have little scientific value. Together, the world’s natural history collections comprise a huge data resource about biodiversity and how it changes.
There are so many specimens at Luomus that no one has a clear picture of the whole. Only a small subset of the specimens have been recoded either to a database or on a paper-based cataloging system. Traditionally, all information about the specimens have been recorded onto small labels attached to them. However, information stored this way is nearly impossible to find, and cannot be analyzed as a whole. For this reason, most of the specimens have not been used after acquisition.
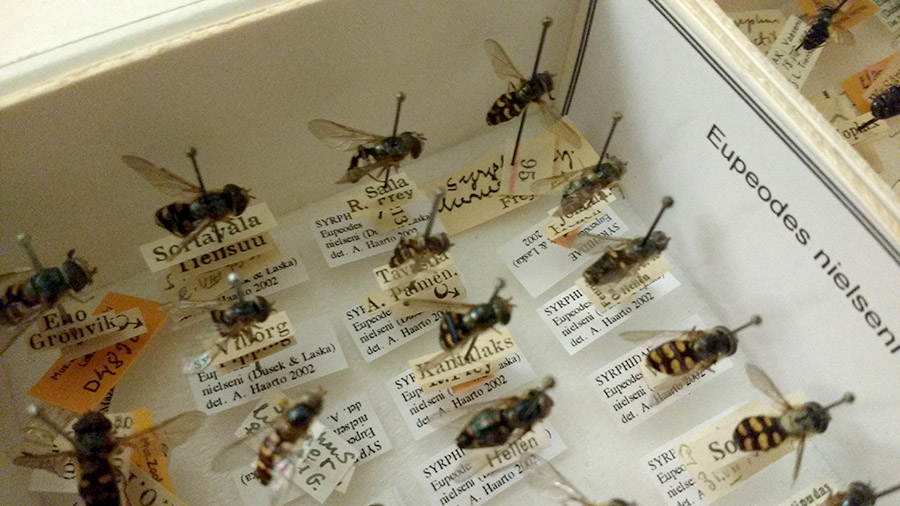
Old insect specimens from the collections of Luomus have labels that have yellowed over time. Later, researchers attached pure white ones to supplement them. Orange labels contain loan codes, meaning that these specimens have been loaned to a researcher at some point.
To address the low availability of the data, projects have been started in the past to digitize the specimens. This development begun at Luomus in the 1960s, when the first specimens were recorded on punch cards. In the 1980s, staff began recording vascular plant specimens into a database. Database creation evolved inconsistently, and by 2000 only a small part of Luomus’ specimens were digitized. At that time, a so-called Luomus Project was started in order to build a system for all biodiversity data. Ambitious plans were made, but the project was eventually shut down because of lack of funding.
From the 1990s onward, many departments at Luomus created their own Access databases, Excel templates and other methods of storing digitized specimen data. These developments were dependent on the work of individual curators and collection managers, so the results were diverse. Based on one survey, there were over 80 different formats for storing biodiversity data. Most of these were only used by a few people and many of them did not work properly. Something had to be done to make the data more accessible and reliably stored.
The First Prototype
In the spring of 2009, I happened to speak with Jyrki Muona, a senior curator at Luomus. He was studying insect specimens from Costa Rica and was excited about their barcode labels, which you could scan and use to get more information about the specimen in a web browser. Seeing that, I had just one thought: I know how to do that! I started to sketch how a new collection management system could work. Eventually, the idea was subsumed by my principal duties, but the excitement persisted.
During the quiet days between Christmas and New Year’s 2009, I set aside some time to program a prototype of a collection management system. I did this independently, without asking anyone’s formal permission. I named it Hyntikka, from the Finnish words for ”very simple specimen database.” Essentially, it was a minimum viable product (MVP), the simplest possible but still useful system. Because of its simplicity it was surely against ”best practices”, but it fulfilled its purpose.
Hyntikka was eagerly put into use, and during 2010 insect department employees at Luomus begun to record specimen data into it. I sought feedback from its users and slowly developed the system further alongside my other duties. At the end of 2010, 23,000 specimens had been recorded into the database. Hyntikka became more and more essential for Luomus, particularly because a database was needed for the data coming from the new digitization center Digitarium.
Development Speeds Up
In the fall of 2010, a new discussion was started at Luomus about the future of its collection management processes, and possibly those of other Finnish museums. We ended up comparing a few alternatives: joining the DINA Project led by The Swedish Museums of Natural History, using commercial Earthcape or open source Specify software, or developing the system on our own. Joining DINA was tempting, but it didn’t feel agile enough with our needs and resources. So in early 2011, we decided to continue our own development, based on our experiences from Hyntikka. The new system was named Kotka (derived from the Finnish words for ”collection database.”)
In addition to specimen data, new needs arose at Luomus to store all kinds of biodiversity information, such as taxonomy (names of and relations between organisms) and nature observations. In order to keep the system simple and manageable by just a few developers, we decided to use an ontology database, a solution that had worked with vascular plant databases since the 1980s. This would be much simpler to build and extend than a traditional relational database (for example, the Specify system has thousands of fields in over 160 tables, whereas ours started with about a dozen fields and a few tables). The decision drew some amazement from those used to traditional databases.
A few other nontraditional and simplified solutions were also brought into use. Approval or authorization processes would be kept very simple. Most of the information would be stored as free text (denormalized), which is well suited to historical data that is rarely modified and also speeds up data recording. Full change histories would be stored (Wikipedia style) in case of errors.
The system would be developed using agile methods using Scrum, adding features one at a time and expanding in cycles of two to three weeks. Use of the system would also be extended into new collections one by one. Problems would be solved when they arose. Development started in the fall of 2011, at which time my role changed from webmaster to lead developer. In practice, I had created a new job for myself.
Agile Maneuvers
The beginning was not without problems. The ontology database was more difficult to take into development use, especially because it was developed slowly alongside other duties. When the delay continued for more than a month, I decided to solve the situation by replacing the ontology database with a MongoDB document database (a “technical pivot” on a Lean startup glossary). After this, I was able to build the user interface. The first version was ready to be tested by real users within a few weeks.
For the next two years, I developed Kotka alongside my other work, publishing new versions periodically and collecting user experiences (iterating!) Hyntikka was also still in use. After spending all of 2013 on a parental leave, I returned to work on Kotka. Now development really took off: A full-time programmer had been hired for the project and I was able to to lead the development as a part-time product owner. Once again, I had a new job description! At the end of 2014, there were about 52,000 specimens in Kotka from the Luomus and Kokkola natural history museums. Kotka was migrated back to the Oracle ontology database, which also included taxonomy data.
Expanding to a National System
In 2014 – in one user’s words – Kotka reached “adulthood”: It worked well for collection management and it was easy for new users to learn. I became a full-time product owner and data migrator. The development efforts were targeted more onto migrating new collections into Kotka and expanding it for botanical gardens. In principle, it was easy to migrate new collections to Kotka (the database and features were very flexible), but cleaning and harmonizing old datasets into a common format was a bottleneck. Nevertheless, new datasets were moved to Kotka from Hyntikka and Digitarium, among others. During 2014, the number of specimens tripled to 150,000.
In 2015, a new Finnish Biodiversity Information Facility (FinBIF) project was started, aiming to gather and share biodiversity information from scattered data providers. Kotka easily became part of FinBIF and expanded to a national service. I started to introduce the system for other Finnish museums. At the end of 2015, seven museums were using Kotka, and some 440,000 specimens had been recorded into it. In the fall 2016 there were 700.000 specimens from nine museums across Finland.
A small side project had grown into a national information system! (For comparison, the Finnish National Museum’s collection database had 220,000 records in 2015.)
So far, Kotka had been developed mostly by just two people, but in November 2015 we were able to hire a third team member to assist in migrating data. Kotka’s reception has been warm; one collection manager commented, “it’s lovely that we can use it!” An important reason behind the quick development of Kotka is that Luomus has its own ICT-team that has specialized in the needs of natural sciences and can make decisions on its own. It’s much easier to work together with end users daily, compared with outsourced programming work, waterfall methods or decisions by an official committee meeting only occasionally.
Update, March 2017: Kotka currently contains over 950,000 specimen records.
Update, September 2022: Kotka currently contains over 3.7 million records, which are available as open data at FinBIF web portal.
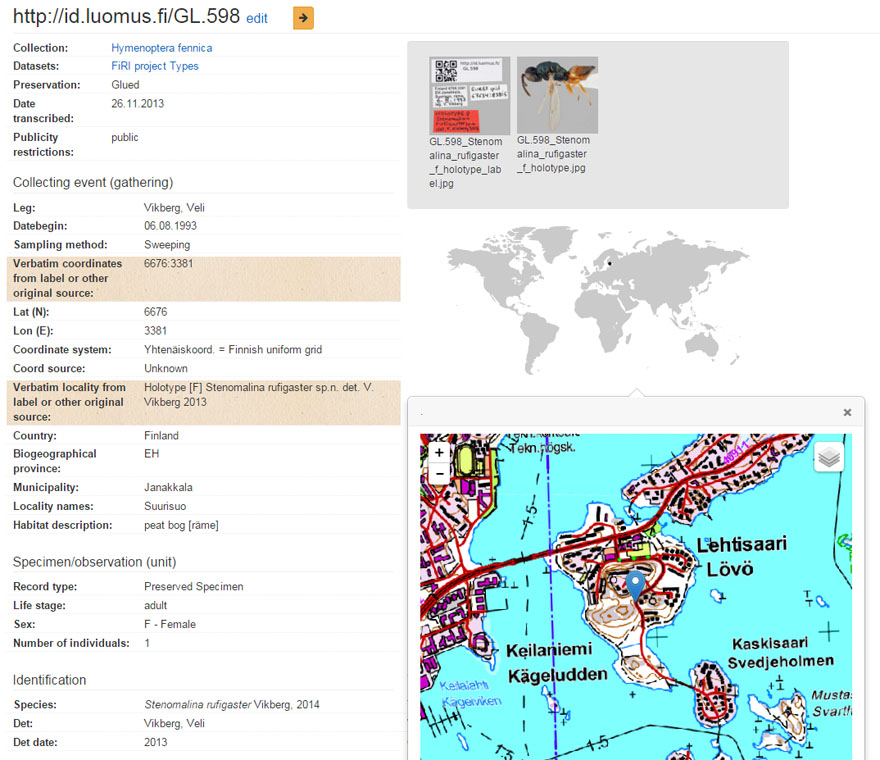
Specimen http://id.luomus.fi/GL.598 data shown on Kotka.
The Future
Based on data volume, the journey of Kotka has just begun. There are still dozens of different databases and sheets, and millions of undigitized specimens. Other challenges also lie ahead: Most of the problems in developing information systems are not technical, but are about communication, understanding different kinds of people and changing the ways people work. How can a mutual understanding about the ultimate goals of the system be achieved and concentrated on, instead just digitizing old and heterogenous work processes? There is still much work to do, but the foundation is now solid. Kotka will evolve onward, based on continous feedback and refinement, as it always has.
Kotka is open source, and available from Bitbucket. It is quite tightly connected to other databases at Luomus (mainly for taxonomy and user management), so it cannot be installed on its own. The ontology database would also take lot of work to install, but there are plans to migrate Kotka to a more generic Oracle JSON database.
